Will nanopores enable single molecule protein sequencers?
Brain storming about nanopore based protein sequencing generates new hypotheses to test

A new perspective in Nature Biotechnology recently highlighted the challenges of developing high throughput protein sequencers.
Sequencing proteins is difficult because the translation of mRNA to protein is the end of the line for our genetic code.
Francis Crick famously wrote in 'On Protein Synthesis' in 1958 that the Central Dogma was "that once ‘information’ has passed into protein it cannot get out again."
This has remained true, biologically, for the last 70+ years, but that's not stopping us from attempting to glean information from proteins via sequencing.
The concept of the central dogma is important when considering how we might go about sequencing proteins, because in the case of RNA and DNA, there exist enzymes we can co-opt to help with performing nucleic acid sequencing.
These include DNA, RNA and RNA dependent DNA polymerases (reverse transcriptases).
But no enzymes currently exist that can "read" a protein and spit out a sequence like theses polymerases do!
So we have to get creative if we want to sequence proteins!
But protein sequencing gets significantly more complicated than nucleic acid sequencing because proteins are built from 20 amino acids, not 4 bases, and each of those 20 amino acids can be modified to create alternative proteoforms!
So, for proteins, there's a lot more signal to deconvolute (and sometimes bonds to break) to get at the sequence.
There have been a number of proposals for how to do this and most of them come down to protein end degradation and recognition, "reverse translation," and direct sequencing with nanopores.
Protein end degradation based sequencing has been around for half a century or more, and you may have heard of "Edman degradation" which is a chemical method to chop amino acids off the ends of proteins.
There are at least two companies (Quantum-Si, Erisyon) working on low to medium throughput peptide sequencers that are based on degradation and recognition of terminal amino acids as peptides are cyclically degraded.
There are also companies (Encodia, Glyphic, AbrusBio) who have combined these degradation schemes with chemistry that "reverse translates" protein sequences into DNA sequences.
This is usually done with DNA oligo tagged antibodies that recognize specific amino acids and when these oligos come in close proximity to one another they're ligated together.
The protein sequence is then determined by sequencing all of these ligated oligos on a DNA sequencer.
But possibly the simplest, in theory, way to go about sequencing proteins is to look at them directly using nanopores!
We've figured out how to do this to sequence DNA and RNA, so the next logical step is to use nanopores to sequence proteins too.
But unlike DNA and RNA which have a strong negative charge, proteins do not have a single charge - regions of them can be positive, negative or neutral!
This makes DNA and RNA relatively easy to pull through a nanopore, but we have to get more creative if we want to use nanopores to sequence proteins.
Some of these creative solutions can be seen in the figure above which reviews some proof-of-concept protein sequencers: a) an unfoldase (unfolds proteins) feeds an unfolded protein into a proteasome (chops up proteins) and peptides are detected as they pass through a pore b) a peptide is linked to a piece of DNA and threaded through a pore c) proteins are end tagged and pulled through a pore d) proteins are put in a special buffer and are pushed through a pore by electro osmotic flow (EOF) e,f) proteins are tagged and then either pulled (e) or pushed (d) through a pore by an unfoldase, and g) an unfoldase feeds a linearized peptide through a pore for sequencing.
All of these methods look interesting, but what I found most exciting about this paper (because I hadn't considered this before!) was how the authors proposed to go about evolving protein sequencing from "identification" to direct sequencing.
This is highlighted in the figure below where in a) they suggest building consensus squiggles (plots created as amino acids block the flow of current while transiting a pore!) for each protein to identify proteins and generate a reference library, b) use that reference library to identify proteins and make suggestions for where modifications might be residing based on changes in peak intensity, c) correlate squiggles to DNA to identify specific proteins, d) and as the technology further improves, identify amino acids and modifications directly.
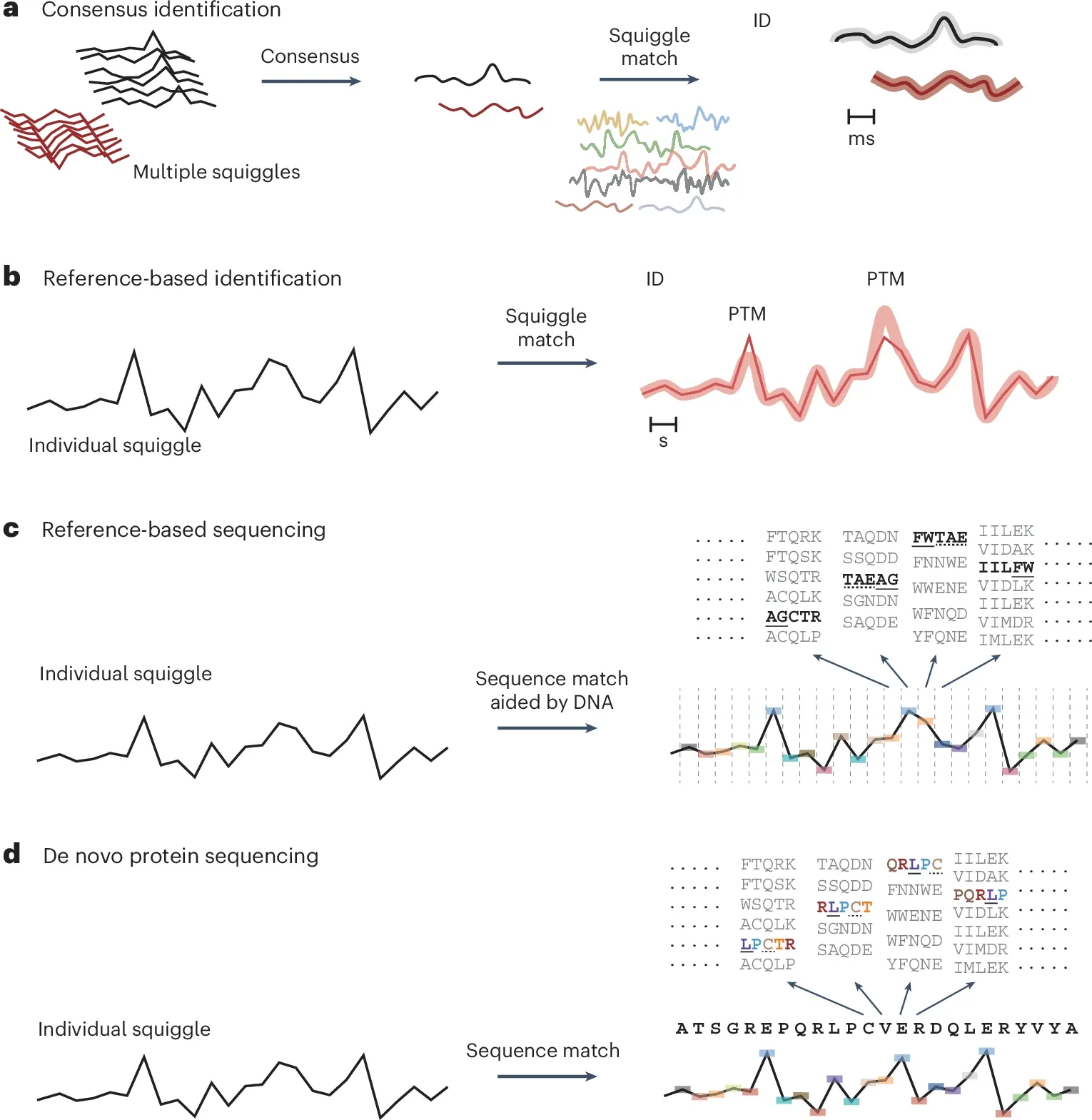
A technological evolution like this means that nanopore based protein detection doesn't have to achieve perfection out of the box and can supplement the other imperfect methods we have now for protein detection like antibody/aptamer immunoarrays and mass spectrometry.
Whether nanopore based protein sequencers will ever be able to address the dynamic range problem (proteins come in varying abundance from one molecule to hundreds of thousands - detecting that wide of a range is hard!) remains to be seen, but 10 years ago some thought that nanopore DNA sequencing would be impossible.
I'm hoping for similar innovations in nanopore protein sequencing that will silence the naysayers once again.
###
Lu C, et al. 2025. Toward single-molecule protein sequencing using nanopores. Nature Biotech. DOI: 10.1038/s41587-025-02587-y
